GENETIC CODE:
CONTENTS:
- Introduction
- Protein synthesis (translation)
- Variations in genetic code
- Wobble
- Universal code
INTRODUCTION:
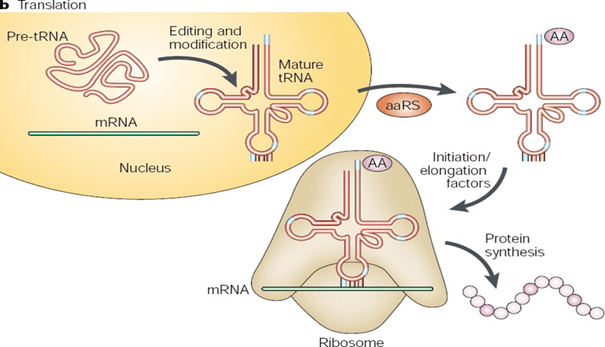
BIOSYNTHESIS OF PROTEINS (TRANSLATION):
The final step in the expression of protein synthesizing genes is translation. The process of Protein synthesis is called translation because it is a decoding process of highly coiled substance. The information encoded in the language of nucleic acids must be written in the languages of amino acid forming proteins. During translation, the sequence of nucleotide is “read” in the appropriate sets of three nucleotide, each set being called a codon. Each codon codes for single amino acid.
- The sequence of codons is “read” in only one way-the reading frame- to give rise to the amino acid sequence of a polypeptide. Deciphering the genetic code was one of the great achievements of the twentieth century.
Close inspection of the code reveals several features that are related not only to the way cells use DNA to store information but also to why it is valuable for storing data.
VARIATIONS:
- One feature is that the code words (codons) are three letters (bases) long; thus one small “word” conveys a significant amount of information. Each codon is re-organised by an anti-codon present on a tRNA molecule.
- Another feature is that the code has “punctuation”. One codon, that is AUG, is always appears as the first codon in the protein synthesizing portion of mRNA molecules in very complicated and important process.
- It is called the start codon because it serves as the start site for translation (a process of protein synthesis) by coding for the internal tRNA. Three other codons (UGA,UAG and UAA) are involved in termination of translation and are called stop or nonsense codons. These codons never encode for an amino acid and therefore they don’t have a tRNA bearing their own anti-codon.
- Thus only 61 out of 64 codons in the code, are known as the sense codons which directs amino acid to incorporate into protein. Finally, the genetic code exhibits code degeneracy; that is, there are up to six different codons for a given amino acid.

- Despite the existence of 61 codons, there are fewer than 61 different tRNAs. It follows that not all codons have a corresponding tRNA.
- Cells can be successfully translated into mRNA using fewer tRNAs because there is not so strong pairing between the 5’ base in the anti-codon of the cell and the 3’ base of the codon which is almost tolerated.
- Thus as long as the appearing first and second bases in the codon correctly base pair with an anti-codon, the tRNA which is bearing the correct amino acid will bind to all mRNA during translation process. This is evident on inspection of the code.
WOBBLE:
The codons for particular amino acids are most often different at the third position. This somewhat loose base pairing which is known as wobble, and it enhance cells the need to synthesize so many tRNAs. Wobble also decreases the effects of particular mutations.
UNIVERSAL CODE:
- The description of the genetic code just provided is of the universal genetic code. However, there are exceptions to the code.
- The first exceptions discovered were stop codons that encoded one of the 20 amino acids. For instance, some protists have a single stop codon (UGA); the other two stop codons are recognized by tRNAs bearing glutamine (Gln).
- More dramatic deviations from the code have also been discovered. Proteins from members of all three domains of life have been discovered to contain the amino acid selenocysteine, the twenty-first amino acid.
- Pyrrolysine, twenty-second amino acid, can be found in the proteins of several methanogens and at least one bacterium.
- Genomic analysis indicates that pyrrolysine might also exist in many other bacteria and some eukaryotes. Selenocysteine is inserted at certain UGA codons, whereas pyrrolysine is inserted at UGA codons.

One thought on “GENETIC CODE”