BY: RAHUL ANDHARIA (MSIWM001)
Introduction:
Genes of an organism is encompassed by Genome. Gene is a hereditary unit, which consists of all the nucleotides and forms a part of chromosome. Spatial distribution of chromatin(DNA and protein) within a cell nucleus is called as Nuclear Organisation. There are different levels in nuclear Organisation.
History:
Chromosome organisation into distinct regions within a cell nucleus was first observed and discovered in the year 1885 by Carl Rabl. With advancement in microscopy technologies, in the year 1909, chromosome territories were discovered by Theodor Boveri when he observed that chromosomes take up separate nuclear regions.
Prokaryotic Organisation:
- Not well defined nucleus is not present in Prokaryotes. Prokaryotes means, ‘before nucleus’. Chromosomal DNAin prokaryotes is present in structures called Nucleoid.
- Circular DNA is present in loops or domains in the nucleoid binded to scaffold proteins which are attached to cell membrane.
- Bacterial chromosomes are huge, but still it gets fitted inside the cell. This occurs because of Supercoiling of DNA.
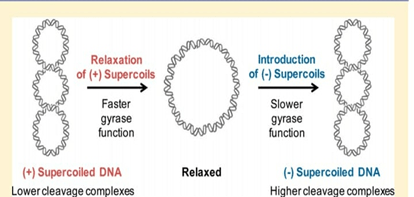
DNA Supercoiling:
- Prokaryotes, compress their DNA into small pieces by supercoiling.
- In bacterial genome the circular DNA is supercoiled because of addition of turns in the double helix structure of DNA.
- Two enzymes are necessary for supercoiling in prokaryotes, Topoisomerases(type 1) and DNA gyrase.
- Chromosomal DNA is compacted to 1000 folds in order to fit DNA within bacterial cells.
- DNA is organised into loops by proteins and further this loops compact the DNA by 10 folds.
- Supercoiling folds DNA by additional 100 folds.
- Supercoiling of DNA can be negative or it can be positive. Negative supercoiling takes place in the opposite direction to the double helix.
- Positive supercoiling is when twisting occurs in the same direction as that of double helix DNA.
During normal growth, most of the bacterial genomes are negatively supercoiled.
Proteins involved in Supercoiling:
- Multiple proteins are involved in DNA folding and condensation, which was reported in the year 1980s and 1990s.
- One of the most abundant protein in nucleoid, HU, binds DNA with the help of enzyme Topoisomerase I, introducing sharp bends in the chromosome, generating tension which is necessary for negative supercoiling.
- One more protein called Integration Host factor(IHF) binds to specific genome sequences and produces additional bends. This was showed by Rice et.al in the year 1996.
- Topoisomerases and DNA gyrase maintains the supercoiling, once the genome gets condensed.
- Genes involved for modulating response to environmental stimuli can be altered in terms of their expression patterns, and this is done by H-NS which is a maintenance protein during transcription.
How these tightly packed genes gets accessed by proteins?
- Nucleoid in Prokaryotes appears as irregular mass, but when cell is treated with chemicals for inhibiting transcription or translation, Nucleoid becomes Spherical.
- Small regions of chromosomes project out during transcription process from nucleus to cytoplasm. In this region they unwind and gets associated with Ribosomes and thus allows easy access for various different types of transcriptional proteins.
Eukaryotic Organisation:
- Large amount of eukaryotic DNA is packed in chromosomes present within nucleus. DNA and proteins together constitutes a chromosome.
- Mostly histone proteins are present in abundance, but certain other proteins are also present in less amount termed as Non- histone proteins.
- DNA-protein nuclear complex is called as chromatin.
- Length of packaging varies, for example in humans 1.6cm shortest DNA molecule can be wrapped and as large as 8.6cm can also be wrapped.
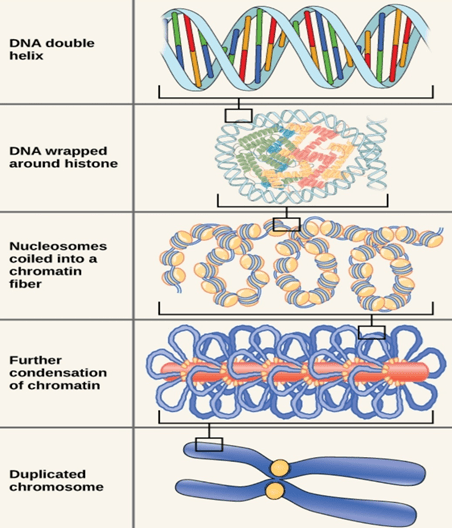
- Packaging of DNA into chromosomes involves folding and has different stages involved in it.
Stage 1(Nucleosome):
- Chromosomal DNA binds to histone proteins in first stage of organisation.
- DNA wraps around histone octamer(2copies of dimers and tetramers each) at regular intervals. DNA-histone complex is called the chromatin.
- DNA wrapping around histone proteins forms bead like structure which is called Nucleosome.
- Nucleosome core particle wrap DNA for about 1.67 left handed turns and contains 146base pairs of wrapped DNA.
- DNA that connects to Nucleosome is called as Linker DNA.
- DNA in this form is seven times shorter when compared to double helix structure without histone proteins. The size of beads is 10nm in diameter compared to double helix which is 2nm.
- Histones are: H1, H2a H2b H3 and H4. H1 is not included in core histone and it connects to linker DNA.
- Histone proteins are basic proteins mostly made up of lysine and arginine amino acids which are positively charged.
Stage 2(30nm fibre):
- In this level of compaction, linker DNA and Nucleosome are coiled into a 30nm fibre.
- Because of this coiling, the chromosome further gets shortened and becomes 50 times more shorter than the extended form.
- Fibre is formed when H1 histone binds to linker DNA at each Nucleosome. When H1 histone interacts with each other, nucleosomes gets pulled together.
Stage 3 (Radial Loops):
- When chromosomes are deprived of histones, they possess a central fibrous protein scaffold, to which DNA gets attached in loops.
- The main role of these fibrous proteins is to make sure that each chromosome in non-dividing cell occupies a particular area of nucleus that doesn’t overlap with other chromosomes.