BY: Reddy Sailaja M (MSIWM030)
A molecular marker is a specific gene fragment present at a specific position called ‘locus’ (pleural loci) in the genome of a cell. These molecular markers are ‘phenotypically neutral’ i.e., they won’t exhibit any genotypic or phenotypic properties. Rather, they ‘flag’ or give ‘sign’ of a particular gene, its functions, variations and inheritance. They act as ‘tags’ if they are present in close proximity of a gene of interest.
These markers help to detect a particular character/trait by analyzing the variations that occur in a particular gene fragment over a period of time.
Ideal characteristics of a molecular marker:
- Polymorphic
- Co-dominant inheritance
- Frequent and even incidence across the genome
- Easy, cost and time effective to use
- Consistency in results
- Reliable data across globe
- Reproducibility
There are three types of molecular markers as follows:
- Morphological markers: These markers are widely used in animal breeding and selection of superior quality farm animals. Morphological characters like skin color, body structure, coat color etc. were considered on visual observation and classify superior quality breeds. However, this technique is not always accurate.
- Cytological markers: These are used to identify proper location of a gene, its genetic diversity with respect to chromosome number and structure in the domesticated animals in comparison to their wild ancestors. Karyotypes, translocations, insertions, deletions, repeats etc. are the characteristics of these cytological markers to be investigated to understand the function of the gene of interest in inheritance, principally in the origin and phylogenetic classification of a species.
- Biochemical markers: Blood type and isozymes are the major biochemical markers that are being investigated at protein level with respect to amino acids composition. These are helpful in understanding phylogenetic relationships at intra and inter-species level. But these are not widely used as proteins are not genetic entities, but an end product of the expressed gene after many modifications.

Figure 1: Types of molecular markers
In short, molecular markers at gene level are the more reliable markers to understand variations and inheritance of a particular gene.
In this session, four major types of molecular markers are discussed as follows:
- Restriction Fragment Length Polymorphism (RFLP): This is one of the early and widely used techniques for DNA analysis. Main principle of the technique is to generate restriction fragments with different sizes that were formed because of nucleotide base insertions, deletions, substitutions, inversions, and duplications etc. in the gene of interest that belongs to the same species. This technique helps scientists to generate gene map/ profile/ finger printing of a particular disease. This technique helps to understand the genetic disease inheritance within the family like hemophilia, a rare blood disorder. Autoradiography (using radioactive probes) or Chemiluminiscence (enzyme linked probe labeling) are the common methods used to visualize the RFLP results.
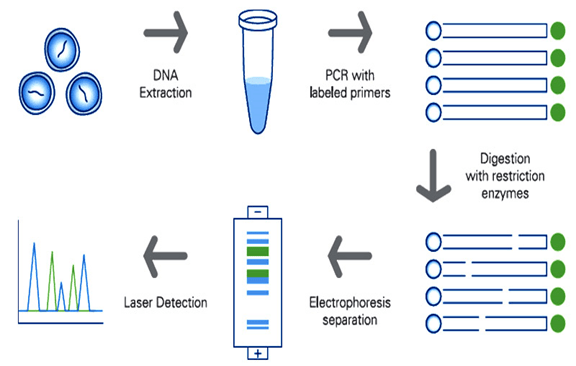
Figure 2: RFLP process illustration
Advantages:
- Simple
- Co-dominant markers expression
- No PCR is requirement
- Distinguish homozygous or heterozygous condition
Disadvantages:
- Needs large amount of pure DNA
- Identification of suitable markers is laborious
- Time consuming
- Requires trained technician to operate
- Random Amplified Polymorphic DNA (RAPD):
RAPD is the most widely used technique to develop DNA markers. It uses short, random oligonucleotide primers (10 – mers) that amplify random sequences at various loci and the PCR to detect variations in the genome. This is a dominant marker selection system and detects polymorphism by analyzing difference in the primer binding site in the DNA sequence between closely arranged sequences of less than 2kb (kilobases).

Figure 3: RAPD process illustration
Advantages:
- Requires small amount of DNA
- Doesn’t require specific primers
- Detects polymorphism effectively
Disadvantages:
- It’s a dominant trait
- Sensitive to PCR conditions
- Generation of non-parental bands in the progeny of known pedigree warns its use
- Not reproducible
- Amplified Fragment Length Polymorphism:
AFLP is a combination of RFLP and polymerase chain reaction (PCR) techniques that detects variation in the DNA sequence from two individuals of a species. Whole genome is digested with the known and rare restriction endonuclease to generate DNA fragments. Adapters are linked to these DNA fragments and primers that are complimentary to these adapters are used to amplify the DNA fragments.
Fragments that are generated by PCR are analyzed using agarose gel electrophoresis. AFLP technique was majorly used in determining genetic variation in the population and has applicability in phenotyping, population genetics, DNA finger printing and quantitative trait loci (QTL) mapping.

Figure 4: AFLP process illustration
Advantages:
- Sensitive – can distinguish homo and heterozygotes
- Wide range of applicability
- Gene mapping
Disadvantages:
- Expensive
- Require large amount of DNA
- Needed trained personnel for sequencing the gels.
- Simple Sequence Repeats (SSR)/Microsatellites:
SSRs are 1 – 6 nucleotides in length and are present throughout the genome as repeats in most of the eukaryotes and a few prokaryotes. They occur as di-, tri-, tetra nucleotide repeats that occur 5-20 times in the genome. The number of repeats varies among different alleles of a gene among population. SSR uses unique sequences as primers that act as flanking regions of a specific DNA fragment. These DNA fragments are further amplified by PCR to generate enough DNA for visualization on agarose or polyacrylamide gels.

Figure 5: SSR process illustration
Advantages:
- Simple to use
- Co-dominant marker
- Map based cloning
- Used to identify genetic distances between population, inbreeds and breeding material during evolution.
Disadvantages:
- Development of proper primers for the satellite region is time consuming and costly.
- Require DNA sequencing
Major applications of molecular markers:
- DNA fingerprinting
- Measure of genetic diversity among the species
- Selection of a Genotype
- Marker assisted selection of a particular trait

One thought on “MOLECULAR MARKERS”