BY- ABHISHEKA G.(MSIWM013)
INTRODUCTION:
1.Recombinant DNA or RDNA technology is defined as the procedure of joining DNA molecules of two different species together and inserted into the host organism to produce a variety of new genetic combinations. This is also known as Genetic engineering.
2. The DNA fragments are selected from two different species and combined. This technique was developed by two scientists namely Boyer and Cohen in 1973.
3. The DNA molecule which is inserted into another DNA molecule is called a VECTOR. The recombinant vector is then introduced into a host cell where it replicates itself, and the new gene is produced. This is the basic principle behind Recombinant DNA technology.
TOOLS OF THE RECOMBINANT DNA TECHNOLOGY:
- Restriction endonucleases: These are used to cut DNA molecules at specific sequences into many smaller DNA fragments.
- Plasmids: These are extrachromosomal circular DNA present in the bacteria, which can replicate independently. During cloning, these plasmids carry drug resistance genes that are used for selection. Foreign DNA can be placed into a plasmid and it is replicated further.
- DNA ligase: This enzyme is used to join the two pieces of DNA together.
- Foreign DNA: This is also known as passenger DNA, which contains desired gene sequences.
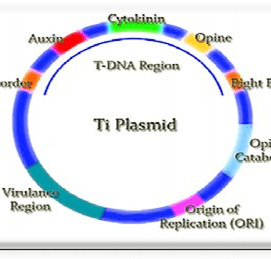
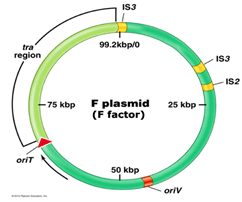
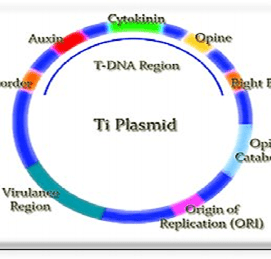
- Vector: It is a vehicle used to insert the desired DNA into the host cell. Some of the vectors used are Plasmid DNA, Bacteriophage DNA, Yeast DNA, Viral DNA, Bacterial DNA, etc.
GOALS OF RDNA TECHNOLOGY:
- To isolate and characterize a gene or DNA from an organism.
- To eliminate undesirable phenotypic characters.
- To combine the needy and beneficial traits of two or more organisms.
- To make desired alterations in one or more isolated genes or DNA
- Inserting the altered genes or DNA into the host cell of another organism.
- To synthesize new genes using artificial methods.
- To alter the genome of the organism
- Understanding the diseases which transmit due to heredity.
- Understanding the treatment for heredity related disorders.
- To create new gene combinations.
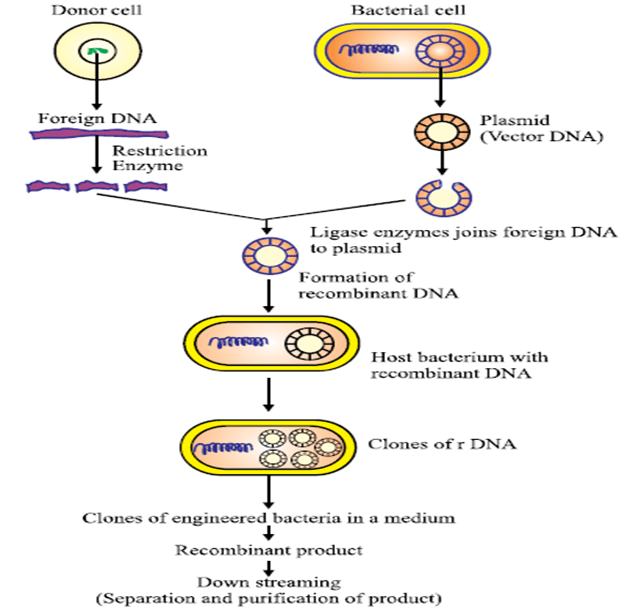
PROCEDURE TO PREPARE RDNA:
1 Isolation of DNA from the organism: The cells are lysed using detergent mixtures, which creates pores in the plasma membrane. Then the mixture of cell contents is treated with protease and RNAase enzymes. The enzyme protease destroys the proteins present in the mixture and the enzyme RNAase destroys the RNA molecules present in the mixture. Then the mixture is centrifuged and the supernatant containing the DNA is transferred into a clean test tube and the DNA precipitated with the addition of ethanol.
2. Insertion of foreign genes into vectors: By using plasmid as a vector, isolated from the bacterial cell and treated with restriction enzymes and target DNA is obtained and it is placed into a vector to produce recombinant DNA.
3. Insertion of Recombinant DNA into host cell: The plasmid containing the foreign DNA is placed into a bacterial or host cell for multiplication.
4.Transformation: The vector is used as a vehicle to transport the gene to host cell, bacterium or other living cells are used as vectors. The vector is multiplied in the host cell and produces many identical copies, which are similar to both DNA and gene present in the DNA.
5. Cloning: After the division of the host cell the rDNA copies produced are transmitted to the progeny and further vector replication takes place in the progeny cell, with the continuous division of cells, a clone of identical host cells is formed. Each clone contains one or more copies of the rDNA molecule. Later the identical host cells are lysis and rDNA molecules are separated from the host cells.

APPLICATIONS OF RDNA TECHNOLOGY:
- This technology helps to grow crops which are resistant diseases and pesticides, crops of our choice, fruits, and flowers of attractive colors.
- This technique is employed in the production of artificial insulin and to deliver the drugs to target sites.
- Used in Molecular diagnosis of diseases.
- Used in Gene therapy.
- Employed in DNA fingerprinting.
- Used in the production of vaccines and pharmaceutical products.
- In the production of monoclonal antibodies.