BY: RAHUL ANDHARIA (MSIWM001)
Introduction:
- In higher eukaryotes, gene expression is tissue-specific. Moderate to high expression of a single gene or only certain cell types show a group of genes. This can be explained through an example, (genes encoding globin proteins are expressed only in erythrocyte precursor cells, called reticulocytes).
- A target gene can be cloned by isolating the mRNA from a specific tissue using this information. Bacteriophage vectors are used to clone specific DNA sequences synthesised from mRNA copies of particular cell type. A fully transcribed mRNA results in cDNA (complementary DNA) which contains only the expressed genes of an organism and replicas (Clones) of such DNA copies of mRNAs are called cDNA clones.
- A combination of cloned cDNA fragments constituting some portion of the transcriptome of an organism is termed as cDNA library, which can be inserted into a number of host cells. Before translation into protein, mRNA is spliced in eukaryotic cells. The DNA synthesized from the spliced mRNA does not have introns or non-coding regions of the gene, so because of this, the protein under expression can be sequenced from the DNA.
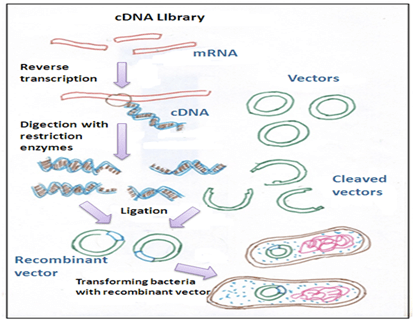
Following steps are involved in the construction of cDNA library:
1. mRNA isolation
2. First and second strand of cDNA synthesis
3. Incorporation of cDNA into a suitable vector
4.Cloning of cDNAs- the cDNA thus obtained is cloned.
Isolation of mRNA:
Total mRNA from a cell type or tissue of interest is isolated. The obtained mRNA is increased in copies by following methods:
- Purification of mRNA by chromatography- in this method oligo-dT column is used which retains mRNA molecules, resulting in their enrichment.
- Density gradient centrifugation method- this method is used to spin down mRNA.
- A string of 50 – 250 adenylate residues (poly A Tail) is present in the 3′ ends of eukaryotic mRNA, which makes the separation easy from the much more prevalent rRNAs, and tRNAs using a column containing oligo-dTs tagged onto its matrix.
- mRNAs bind to the column due to the complementary base-pairing between poly (A) tail and oligo-dT when an cell extract is passed through oligo-dT column. In an unbound fraction ribosomal RNAs and transfer, RNAs flow through. A low-salt buffer is used to elute the bound mRNAs.
First and second strand of cDNA synthesis:
- As, mRNA is single-stranded it cannot be cloned as such and is not a substrate for DNA ligase. It is first converted into DNA before insertion into a suitable vector, which can be achieved using reverse transcriptase (RNA-dependent DNA polymerase or RTase), obtained from avian myeloblastosis virus (AMV).
- Annealing of a short oligo (dT) primer to the Poly (A) tail on the mRNA takes place.
- Enzyme, (Reverse transcriptase) extends the 3´-end of the primer using mRNA molecule as a template producing a cDNA: mRNA hybrid.
- By RNase H or Alkaline hydrolysis mRNA from the cDNA: mRNA hybrid can be removed to give a ss-cDNA molecule.

Cloning of cDNAs:
Vectors generally used to clone cDNAs are phage insertion vectors. The advantages of using Bacteriophage vectors over plasmid vectors are as follows;
- When a large number of recombinants are, required bacteriphage vectors are more suitable for cloning low-abundant mRNAs as recombinant phages are produced by in vitro packaging.
- As compared to the bacterial colonies carrying plasmids, bacteriophage vectors can easily store and handle large numbers of phage clones. Particularly in the isolation of the desired cDNA sequence involving the screening of a relatively small number of clones, plasmid vectors are used extensively.
Applications of cDNA libraries/cloning:
- To discover novel genes.
- Gene functions in vitro studies by cloning full-length cDNA.
- To determine alternate splicing in various cell types/tissues.
- Various non-coding regions from the library can be removed with the help of cDNA libraries.
- For detection of the clone or the polypeptide product, gene expression is required and is the primary objective of cloning.
- To study the expression of mRNA.
Disadvantages of cDNA libraries:
- Parts of genes found in mature mRNA are present in cDNA libraries.For example, (those involved in the regulation of gene expression), will not occur in a cDNA library which are sequences before and after the gene.
- For isolating the genes expressed at low levels, cDNA library cannot be used, as there will be very little mRNA for it in any cell type.
Applications and uses of cDNA library:
Due to the removal of non-coding regions, storage of reduced amount of information can be possible.
- In prokaryotic organisms, cDNA can be directly expressed
- In reverse genetics, cDNA libraries are useful, where the additional genomic information is of less use.
- Gene coding for particular mRNA can be isolated using cDNA library.
cDNA library and Genomic DNA library differences:
- Non-coding and regulatory elements found in genomic DNA are absent in cDNA library.
- Detailed information about the organism can be obtained using genomic libraries, but are more resource-intensive to generate and maintain.
Construction and applications of genomic library
Introduction:
- An organism specific collection of DNA covering the entire genome of an organism is considered as a genomic library. DNA sequences such as expressed genes, non-expressed genes, exons and introns, promoter and terminator regions and intervening DNA sequences are present in a genomic library.
- Some common steps in construction of a genomic DNA library includes; purification and fragmentation of genomic DNA, isolation of gDNA followed by cloning of the fragmented DNA using suitable vectors. Protease and organic (phenol-chloroform) extraction is used to digest eukaryotic cell nuclei. Thus, the genomic DNA obtained is too large to incorporate into a vector and needs to be broken into desirable size fragments. Physical methods and enzymatic methods are used for fragmenting DNA. Representative copies of all DNA fragments present within the genome can be collectively obtained in a genomic library.
Mechanisms for cleaving DNA
(a) Physical method
In this method, genomic DNA is sheared mechanically using a narrow-gauge syringe needle or sonication to break up the DNA into suitable size fragments that can be cloned. About 20 kb fragment is desirable for cloning into Lambda based vectors. Variable sized DNA fragments may result due to random DNA fragmentation. Large quantities of DNA are required in this method.
b) Enzymatic method
- In this method, restriction enzymes are used for the fragmentation of purified DNA.
- The action of restriction enzymes will generate shorter DNA fragments than the desired size and hence this method is limited by probability distribution.
- Multiple recognition sites for a particular restriction enzyme are present in a gene to be cloned; the complete digestion will generate fragments that are generally too small to clone and hence the gene may not be represented within a library.
- Partial digestion of the DNA molecule is usually carried out using known quantity of restriction enzyme to obtain fragments of ideal size, to overcome the problem of multiple sites.
- Type of ends (blunt or sticky) are the two factors that governs the selection of restriction enzymes to be used and are generated by the enzyme action and susceptibility of the enzyme to modification of bases like methylation (chemical modification) can inhibit the enzyme activity.
- Agarose gel electrophoresis or sucrose gradient technique is used to generate fragments of desired size and then ligated using suitable vectors.
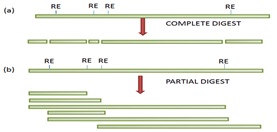
(Figure –The complete (a) and partial (b) digestion of a DNA fragment using restriction).
Cloning of genomic DNA:
There are different types of vectors available for cloning large DNA fragments. λ phage, yeast artificial chromosome, bacterial artificial chromosome etc, are suitable vectors for larger DNA and λreplacement vectors like λDASH and EMBL3 are preferred for construction of genomic DNA library. Selected DNA sequence into the vector is ligated by using T4 DNA ligase.
Advantages of genomic libraries:
- It is used to identify a clone encoding a particular gene of interest.
- Prokaryotic organisms having relatively small genomes can be mapped using genomic libraries.
- Genome sequence of a particular gene, including its regulatory sequences and pattern of introns and exons can be studied from Genomic libraries from eukaryotic organisms.


Hello! I know this is somewhat off topic but I was wondering if you knew where I could get a captcha plugin for my comment form? I’m using the same blog platform as yours and I’m having trouble finding one? Thanks a lot!
LikeLike