BY: SAI MANOGNA (MSIWM014)
Every time a cell divides, DNA breaks each of its double strands into two single strands. Each of these single strands functions as a template for a new complementary DNA strand. Each new cell, as a result, has its complete genome. This method is known as the replication of DNA. Replication is regulated by the pairing of template chain bases with incoming deoxynucleoside triphosphates and is driven by DNA polymerase enzymes. It is a complicated process involving an array of enzymes, particularly in eukaryotes.
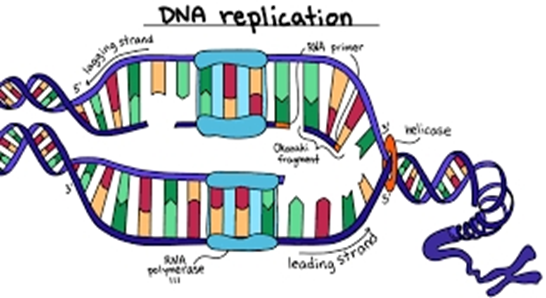
1. DNA biosynthesis begins in the direction of 5′- to 3′. This makes it difficult for both strands to be simultaneously synthesized by DNA polymerases. It is first necessary to unwind a portion of the double helix, and helicase enzymes mediate this.
2. The leading strand is continuously synthesized, but in short bursts of about 1000 bases, the opposite strand is copied as the lagging strand template becomes available. The resulting short strands are called Okazaki fragments.
3. There are at least three distinct DNA polymerases in bacteria: Pol I, Pol II, and Pol III; Pol III is primarily involved in the elongation of the chain.
4. However, DNA polymerases may not initiate de novo DNA synthesis, but require a short primer with a free 3′-hydroxyl group. This is provided by an RNA polymerase (also called DNA primase) in the lagging strand that can use the DNA template and synthesize about 20 bases in length for a short piece of RNA.
5. Pol III can then take over one of the short RNA fragments previously synthesized. Pol I take over at this stage, using it’s 5′- to 3′-exonuclease activity to digest the RNA and fill the DNA gap until it reaches a continuous DNA stretch.
6. This leaves a gap between the newly synthesized DNA 3′-end and the DNA 5′-end previously synthesized by Pol III.
7. DNA ligase, an enzyme that creates a covalent bond between a 5′-phosphate and a 3′-hydroxyl group, aims to fill the gap.

Transcription:
The mechanism of producing an RNA copy of a gene sequence is transcription. This version, called a molecule of messenger RNA (mRNA), leaves the cell’s nucleus and enters the cytoplasm, guiding the protein synthesis that encodes.
One of the fundamental processes that happen to our genome is transcription. It’s the mechanism by which DNA is converted into RNA. And you may have heard of the Central dogma of Molecular biology in which DNA is transcribed to RNA and translated to protein. Well, the first part of moving from DNA to RNA relates to transcription. And in unique locations, we transcribe DNA to RNA. But there are many other transcribed RNAs, including transfer RNAs and ribosomal RNAs, which do other genomic functions.
Formation Of pre-messenger RNA:
Initiation: There are similarities between the transcription machinery and that of DNA replication. As with DNA replication, before transcription can occur, a partial unwinding of the double helix must happen, and it is the RNA polymerase enzymes that catalyze this process.
Elongation: Unlike DNA replication, only one strand is transcribed. The strand containing the gene is called the sense strand, while the antisense strand is the complementary strand. A copy of the sense strand is the mRNA produced in transcription but transcribed as the antisense strand.
Termination: Ribonucleoside triphosphates (NTPs), with Watson-Crick base pairing (A pairs with U), align along the antisense DNA strand. For the formation of a pre-m-RNA molecule that is complementary to a region of the antisense DNA strand, RNA polymerase binds the ribonucleotides together. Transcription stops when a triplet of bases read as a “stop” signal enters the RNA polymerase enzyme.

RNA Splicing: Gene expression is regulated at several different stages during transcription and translation to ensure that the right products are produced. The gene includes different sequences in eukaryotes that do not code for protein. The transcription of DNA generates pre-mRNA in these cells. These pre-mRNA transcripts often contain regions known as introns, which are interfering sequences removed by the splicing process before translation. Exons are called the areas of RNA that code for protein. In a process called alternative splicing, splicing can be regulated so that various mRNAs can contain or lack exons. Alternative splicing makes it possible to create more than one protein from a gene. It is a significant regulatory step in deciding which functional proteins are produced from the expression of genes.

Alternative splicing:
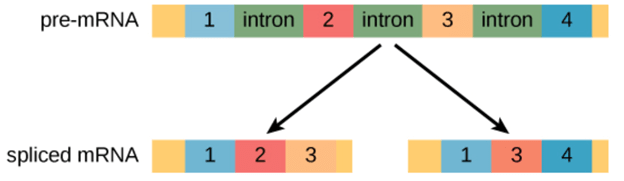
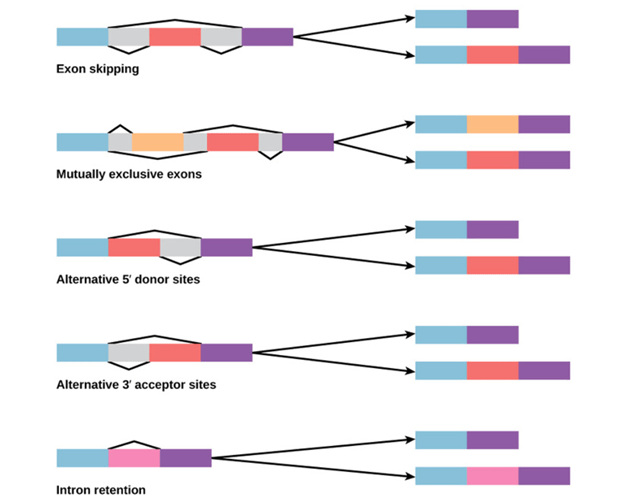
Alternative splicing occurs during gene expression and enables multiple proteins (protein isoforms) generated from a single gene coding process. Due to the distinct forms in which an exon can be exempted from or included in the messenger RNA, alternate splicing can occur. It can also occur if parts of an exon are excluded or included, or if introns are included. For example, if four exons (1,2,3 and 4) are present in a pre-mRNA, they can be spliced and translated into many different combinations. It is possible to translate Exons 1, 2, and 3 together or translate Exons 1, 3, and 4 together.
By binding regulatory proteins (trans-acting proteins that contain the genes) to cis-acting sites located on the pre-RNA, the pattern of splicing and production of alternative-spliced messenger RNA is regulated. Splicing activators (that encourage specific splicing sites) and splicing repressors (that minimize the use of particular sites) are among some of these regulatory proteins.
Heterogeneous Nuclear Ribonucleoprotein (hnRNP) and Polypyrimidine Tract binding protein (PTB) include some common splicing repressors. The proteins translated from; spliced messenger RNAs, alternatively differ in their amino acid sequence, resulting in the protein’s altered function. This is one of the reasons, the human genome can encode a broad range of proteins. A typical process in eukaryotes is alternative splicing; most of the multi-exonic genes in humans are alternatively spliced. Unfortunately, the explanation that there are many hereditary diseases and disorders is often irregular differences in splicing.
Spliceosome:
Messenger RNA splicing is achieved and catalyzed by a macro-molecule complex called the spliceosome. The ligation and cleavage regions are determined by several subunits of the spliceosome, including branch sites and splice sites of 5′ and 3′. Interactions between these sub-units and small nuclear ribonucleoproteins (snRNP) found in spliceosome produce a complex that helps decide which introns to leave out, exons that hold together to bind together. After the introns are cleaved and detached, a phosphodiester bond binds the exons together.
Reverse Transcription: Reverse transcription is a technique used to create a complementary DNA strand from RNA. This mechanism is based on a retroviral mechanism whereby reverse-transcriptase enzymes can reverse transcribe RNA into DNA. This is incredibly helpful when scientists only have tissue and want to study gene sequences. Researchers will separate mRNA from the tissue; in this case, use reverse transcription to generate cDNA.

Translation:
The translation is the process in which proteins are generated using the information carried in mRNA molecules. The nucleotide sequence in the mRNA molecule provides the code for an essential amino acid sequence to create a protein. A protein is made from many amino acids, similar to how RNA is constructed from many nucleotides. A ‘polypeptide chain’ is considered a chain of amino acids, and a polypeptide chain bends and folds on itself to form a protein.
The RNA strand information is translated from RNA’s language into the language of polypeptides during translation, i.e., the nucleotide sequence is translated into an amino acid sequence.

Initiation:
The smaller and larger subunits of the ribosome bind to the mRNA transcript at its binding site during the translation, in the initiation stage.
When the starting codon AUG is recognized by the tRNA, the process of protein construction begins.
Elongation:
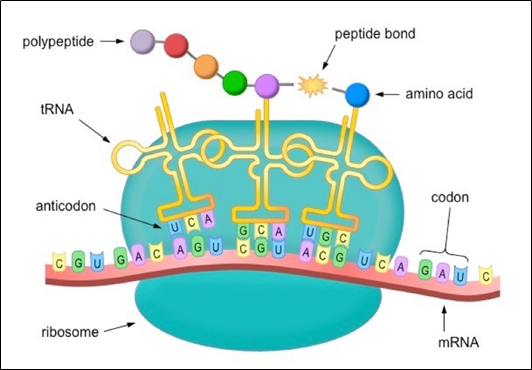
The mRNA triplet codon is “read” during the translation elongation process, and the tRNA is added to the complementary amino acid. Ribosomal RNA catalyzes the whole reaction.
Termination:
If the termination codon is reached, the polypeptide chain synthesis ends with peptidyl tRNA. Here, the whole process depends on the RNA polymerase’s involvement in the transcription, although no polymerase is involved in the translation. Interestingly, transcription is a method of encoding information in mRNA (messenger RNA), while translation is a decoding method.
In eukaryotes transcription occurs in the nucleus, while translation in the cytoplasm. However, the entire process occurs only in the cytoplasm in the prokaryotes.
Rifampicin antibiotics inhibit transcription, while translation is hindered by puromycin and anisomycin. The final transcription product consists of Adenine, Guanine, cytosine, and Uracil messenger RNA. At the leading site, it has the initial codon and, in the end, the termination codon. Due to a long adenine chain, the 3 ‘end of the mRNA is called a poly-A tail. The final translation result is a long amino acid chain, a fundamental building block of a protein called the polypeptide chain. The first amino acid is methionine in the amino acid chain. (Although, in most cases, it is removed).
Transfer RNA (tRNA):
1. Ribonucleic acid transfer (tRNA) is a type of RNA molecule that helps decode a sequence of messenger RNA (mRNA) into a protein.
2. During translation, tRNAs act at specific sites in the ribosome, which is a mechanism that synthesizes a protein from an mRNA molecule.
3. Proteins are formed from smaller units called amino acids, which are specified by codons called three-nucleotide mRNA sequences.
4. A codon represents a specific amino acid, and a particular tRNA is known for each codon.
5. With three hairpin loops forming a three-leafed clover, the tRNA molecule has a distinctive folded structure.
6. A series called the anticodon includes one of these hairpin loops, which can identify and decode an mRNA codon. Each tRNA has attached its corresponding amino acid to its end.
7. The tRNA transfers the necessary amino acid to the end of the increased amino acid chain when a tRNA recognizes and binds to its respective codon in the ribosome.
8. Then, the mRNA molecule begins to decipher the tRNAs and ribosomes until the whole sequence is converted into a protein.

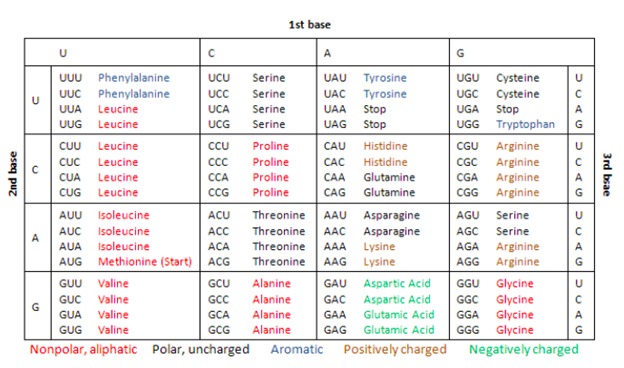
Genetic Code: In the early 1960s, American biochemists Marshall W. Nirenberg, Robert W. Holley, and Har Gobind Khorana completed the deciphering of the genetic code. Genetic code is the term we use for the context in which the four DNA bases — A, C, G, and Ts — are linked together to be read and converted into a protein by the cellular machinery, the ribosome. Each of the three nucleotides in a row counts as a triplet in genetic code and codes for a single amino acid. So, each of the three sequences codes for an amino acid. And often, proteins are made up of hundreds of amino acids. Thus, the code that would make one protein could contain hundreds, sometimes even thousands, of triplets.