BY- K. Sai Manogna (MSIWM014)
Introduction :
Poxviruses are part of the family Poxviridae and can infect humans and animals alike. Smallpox (variola), vaccinia, cowpox, monkeypox, buffalopox, aracatuba, and cantagalo viruses are orthopoxviruses. Orf virus, pseudo cowpox virus, deer poxvirus, bovine papular stomatitis virus, and sealpox virus are parapoxviruses. Yatapoxviruses include the tanapox virus, which is commonly found in Africa. The human poxvirus, the molluscum contagiosum virus, contains mollusc poxviruses.
Smallpox is unique to humans and molluscum contagiosum. In humans, other viruses cause unusual zoonotic infections. The vaccinia virus used for vaccines will infect humans as well. Molluscum contagiosum is also a human-unique poxvirus. Other human poxvirus infections are caused either by zoonotic exposure to animal poxviruses or scheduled or unintended vaccinia administration. This virus is spread through close contact, often via sexual contact.
Recently, laboratory exposures have been reported that have contributed to infection with vaccinia and tanapox viruses, widely used as vectors for experimental vaccines. Due to transmission to contacts, civilian and military personnel’s smallpox vaccination program resulted in multiple infections.
Incubation Period: Infections with smallpox are caused by inhalational penetration to nasal, oral, or pharyngeal droplets. The incubation period is from 10-14. Viruses of smallpox multiply locally and spread to the nearby lymph nodes. On days 3-4, asymptomatic viremia ensues, spreading to the bone marrow and spleen. On approximately day 8, secondary viremia begins. Generalized signs of fever and a toxic appearance are associated with this secondary viremia. In the dermis blood vessels, the virus in leukocytes then becomes localized. Then the stereotypical smallpox rash emerges.
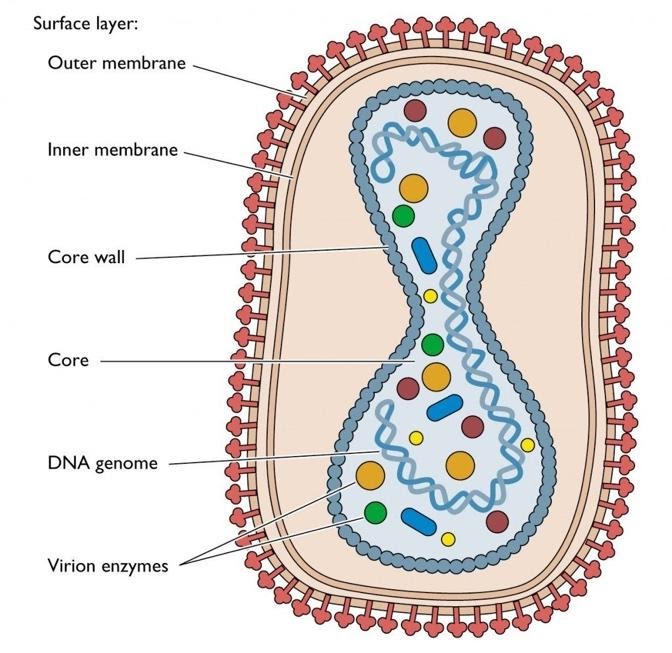
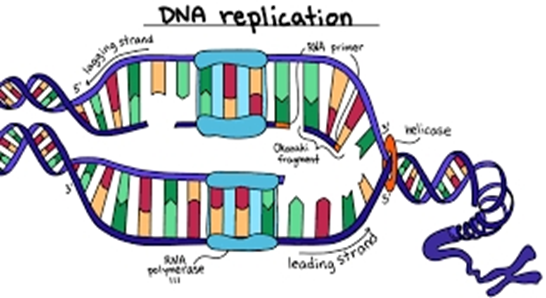
The replication:
1. Generally, poxviruses replicate only in the cytoplasm and can in cells without a nucleus, unlike most DNA viruses.
2. Either through endocytosis or by a fusion event, the virion enters the cell.
3. For later replication cases, the viral core reaches the cytoplasm and functions as a scaffolding.
4. Besides, many necessary enzymes such as viral transcriptase, transcription factors, capping and methylating enzymes, and a poly(A) polymerase are transported by the virus.
5. Therefore viral DNA transcription is rapidly initiated and about 100 early viral genes, especially enzyme coding genes that are involved in viral DNA replication, are triggered.
6. At the same time as DNA replication, transcription of intermediate and late genes is initiated.
7. In specific cytoplasm areas, virus assembly and immature viruses can be visualized very quickly.
8. The Virions travel to the Golgi complex during maturation, where they are enveloped before being released by budding or cell disruption.
9. Some virions are enveloped, and they may have certain benefits, including cell uptake speed.
Other poxviruses, primarily localized diseases, typically follow the same pattern of evolution. An exception is the outbreak of monkeypox, which progresses to a variola-like clinical syndrome. Monkeypox infections, as in the North American epidemic, can vary from mild infections with few lesions to severe systemic diseases that mimic smallpox. At the inoculation site, the Molluscum contagiosum virus also replicates, but the skin lesions’ character is distinct.
Epidemiology:
Human poxvirus infections are obtained from animal reservoirs, except molluscum, which is primarily a human disease. The reservoir is recognized and distributed globally in some cases, as is the case with ovine and bovine parapoxviruses. An occupational hazard of those who interact with the contaminated reservoir hosts is human contamination with these viruses.
Monkeypox is confined to West Africa, and squirrels are more critical than monkeys as reservoir hosts. The cowpox virus is limited to Europe and the western part of the former Soviet Union. Bovine cowpox is unusual, and the most frequently recorded host is the domestic cat. Lack of definitive knowledge on the cowpox virus reservoir host, but it is presumably tiny wild rodents. Cases occur without established contact with cats or cattle, and it is possible to spread indirectly through barbed wire or brambles.
Restricted natural person-to-person transmission of monkeypox, but not more than four or five generations, has been observed. Parapox and cowpox diseases spread from person to person occasionally, if ever. Molluscum’s person-to-person spread is historically associated with physical contact sports (e.g., wrestling) and towel sharing. However, there is growing evidence that molluscum sexual transmission is significant.
Traditionally, the Vaccinia virus is known as a laboratory virus without a natural reservoir. Although its host reservoir lacks information, the buffalopox virus, now considered a vaccine virus variant, appears to have established itself in India. It is important to note that such strains may become developed in animal populations and interact with genetically related viruses circulating in them due to the possible use of recombinant vaccinia virus vaccines.
Diagnosis:
In some instances, a right diagnosis would be made possible by the existence of the lesions and a careful history of interaction with the infected reservoir animal or other infected people; difficulties can occur if no such contact is known. This is perhaps most prominent with human cowpox, as most cases are not linked to a single source, and occasionally an anthrax clinical diagnosis is made.
An effective means of rapid diagnosis is electron microscopy of the vesicle or scab material; poxviruses and herpesviruses are readily differentiated, and the characteristic morphology of parapoxviruses can be identified. Immunofluorescence of infected cell cultures can distinguish morphologically related poxviruses from various genera for e.g., Orthopoxviruses and Yatapoxviruses. While the molluscum virus has yet to be cultivated in tissue culture and chicken embryos, the other poxviruses are easily isolated. Cultivation then enables biologic and serum neutralization samples to be established. Near antigenic relationships within genera compromise precise recognition through antibody detection, but knowledge of the host and geographic range will validate a presumptive diagnosis.
Treatment:
Infections of Variola have been globally eradicated. There is a concern, however, about the reintroduction of smallpox through bioterrorism. An international health care emergency would precipitate the reappearance of smallpox. Local and federal public health authorities should be informed of suspected cases of smallpox.
For smallpox or vaccinia, no known remedies are currently available. Several nucleoside and nucleotide analogues, including variola, vaccinia, monkeypox, cowpox, molluscum contagiosum, and orf, have shown Invitro activity and in vivo antiviral activity against various poxviruses. So far, Cidofovir and a number of its derivatives are the most efficient.