BY: Reddy Sailaja M (MSIWM030)
INTRODUCTION
Infectious diseases affect living forms on earth that include but not limited to plants, animals including human beings. Pathogenic micro organisms i.e., bacteria, virus, parasites etc attack, hinder the growth and development of an organisms and sometimes lead to death.
Infectious diseases that turn out to be pandemic have had bad effect on human beings in the history like bubonic plague, influenza, Spanish flu, avian flu and the most recent on COVID-19 (SARD-CoV-2). When the diseases spread rapidly and cross the country’s border, it is called as a pandemic.
Rapid diagnosis and treatment are the only mode of spreading the disease and to save lives. Standard traditional methods of microbial detection include – microbial culture (aerobic and anaerobic), Gram staining, colony morphology and other biochemical analysis. However, these traditional microbial methods take 5-7 days to give result, by then patients would be severely affected and sometimes might die.
Henceforth, rapid diagnosis with in a day or in hours of time is critical for effective treatment and patient management. Recently, Nucleic acid amplification technique (NAAT), a major molecular biology application has gained interest in the diagnostic field for its rapid and sensitive pathogen detection in short time. Polymerase chain reaction (PCR), a NAAT method gave hope for the early infection detection as it is fast and sensitive method in pathogen detection. PCR uses thermoresistant DNA polymerase enzyme (Taq polymerase), sequence specific primers and under specific cyclic conditions amplify the pathogenic DNA isolated from the sample to billions of copies in few hours. Due to sensitivity and speed, PCR became the choice of pathogen detection in medical microbiology field.
However, PCR has certain draw backs as follows:
- Need of a thermocycler (high cost)
- Need of carcinogenic material like Ethidium bromide for DNA band visualization
- Need of a trained technician
- Sophisticated molecular biology lab setup with at least three isolated rooms
- Can’t be setup at point of care centres like rural areas
Loop-mediated Isothermal amplification (LAMP) is a revolutionary NAAT method, discovered by Notomi et al in 2000. It has the potential to rapidly detect the pathogenic DNA more sensitively and specifically in comparison with PCR at a constant temperature (isothermal). LAMP method is based on the auto cycling of the DNA using strand displacement reaction and utilizes DNA polymerase like Bst, Bsm, Gsp etc
Due to its high sensitivity, speed and efficiency, it has varied applications in medical microbiology field.
| CHARACTERISTICS | LAMP | PCR |
| Type of reaction | Isothermal | Cyclic |
| Reaction time | 30-60 minutes | 2-4 hours |
| Temperature | Isothermal Temperature (60-65⁰C range). | Variable Temperature. Denaturation (95⁰C); Annealing (50-60⁰C); Extension (72⁰C) |
| Need of thermocycler | No. simple dry bath is enough | Yes |
| Sensitivity | Limit of detection is higher. | Limit of detection is lower. |
| Specificity | High as it uses 4-6 primers | Lower than LAMP |
| Sample kind | Detection is good even with crude samples | Pure DNA is required |
| Primers characteristics | 4-6 primers. Loop primers increase reaction speed | 2 primers |
| Mode of detection | Fluorescence detection in real time. Visual detection with naked yes, gel electrophoresis or turbidity. | Fluorescence detection in real time. Visual detection is only through gel electrophoresis |
Table 1: Differences between LAMP and PCR techniques
LAMP PRIMERS
LAMP makes use of 4 primers that are designed specifically to recognise 6 different regions on the target gene.
- Forward inner primer (FIP) – FIP comprises of two primers namely, F2 region at the 3’ end and F1C region at 5’ end (F1C of the primer is similar to F1C portion of the target DNA).
- Backward inner primer (BIP) – BIP comprises of two primers namely, B2 region at the 3’ end and B1C region at 5’ end (B1C of the primer is similar to B1C portion of the target DNA).
- Forward outer primer (F3) – F3 is the outer primers and is short in its length. F3 is complementary to F3C region of the target DNA.
- Backward outer primer (B3) – B3 is one of the outer primers and is complementary to B3C region on target DNA.
- Loop primers – Loop forward and loop backward (LF and LB) are the two additional primers utilized in the LAMP reaction to increase the speed.

Figure 1: LAMP primers
STAGES IN THE LAMP REACTION
1. F2 portion of FIP primer binds to F2C region on the target DNA and initiates new strand synthesis and amplification.
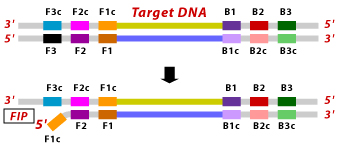
2 F3 outer primer then binds to the F3c region on the target DNA and extends the strand by displacing the FIP associated complementary strand. This displaced strand forms a loop lie structure at the 5′ end.

3. Th synthesized ssDNA with a loop at the 5′ end assists as a template for BIP. B2 binds to B2c region on the target DNA and synthesizes new complementary strand by opening of the 5′ end loop.
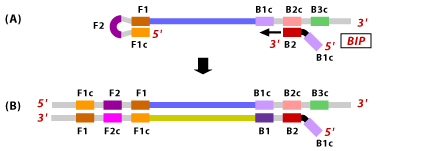
4. Now, the B3 primer binds to B3c region on the target DNA and extends by displacing the BIP connected complementary strand. This results in dumbbell shaped DNA formation.
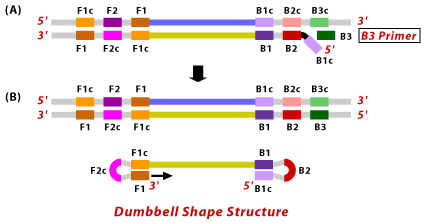
5. Then the F1 primer gets extended by opening up the loop at the 5′ end with the help of Bst DNA polymerase. At this stage, dumbbell shaped DNA become a stem loop structure and initiates the LAMP reaction. This stage is called the LAMP reaction’s second stage.

6. Further in LAMP cycling, the FIP binds to the loop region of the stem-loop DNA and initiates strand synthesis by displacing F1 primer and formation of a new loop at the 3′ end.

7. Extension happens at the 3′ end of B1 by displacing the FIP strand, forming a dumbbell shaped DNA. Self-primed DNA synthesis by strand displacement gives out one complementary strand of the original stem loop DNA along with one more stem loop DNA with a gap repair.

8. Both the new stem loop DNAs act as template for a BIP primed strand displacement reaction in the succeeding cycles. Consequently, for every half LAMP cycle, 13 fold amplification of the target DNA occurs.

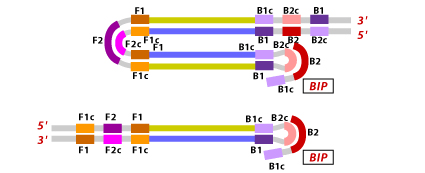

The ultimate amplification LAMP products contain combination of stem loop DNA with varied lengths looking like a cauliflower like structure with multiple loops.
LAMP REACTION SET UP
To setup LAMP assay, target DNA, an isothermal DNA polymerase with strand displacement activity, primers and buffer are sufficient. LAMP assay can be setup in a simple water bath or a heat block at a constant temperature (ideally at 65°C).

Figure 2: Over view of LAMP reaction
LAMP DETECTION
LAMP reaction can be detected as follows:
- Fluorescence based detection: using florescent dyes like SYBR green, Pico green, Eva green

Figure 3: LAMP detection using SYBR green fluorescent dye
- Visual detection: Based on turbidity and precipitation in the positively reacted tubes. Leuco crystal violet is a dye that detects positive reaction based on turbidity.

Figure 4: LAMP detection based on turbidometry
- Colorimetric dyes are also used as they react with the free Mg2+ being produced in the reaction. Colour change happens based on the pH change. For example, phenol red show pink colour before the reaction and turns yellow if the sample is positive for the pathogen (as the pathogenic DNA multiplies, more Mg2+ will be released into the reaction tube).

Figure 5: Colorimetric detection of LAMP reaction (Source: NEB)
ADVANTAGES OF LAMP TECHNIQUE
- Require simple water bath or heat block (no costly thermocycler).
- Amplification at isothermal conditions.
- More specificity and sensitivity as it utilizes 4-6 primers.
- Cost effective
- Easy deployment at point of care centres at rural areas.
- No trained technician is required. Setup is quite simple.
APPLICATIONS OF LAMP TECHNIQUE
- Rapid diagnosis of bacterial, viral, fungal and parasitic organisms.
- Helps to detect pathogens at both genus level and species level. In case of viruses, various strains can be easily differentiated.
