BY- SREELAKSHMI (MSIWM012)
Carbohydrates are polyhydroxy aldehydes or ketones, or substances that produce such substances in hydrolysis. Most, but not all, carbohydrates have a positive effect (CH2O) n.
Stereo Isomerism:
The presence of carbon asymmetric atoms allows the formation of isomers. Chemicals that have the same structure but differ only in localization are called stereoisomers or geometric Glucose isomers with 4 asymmetric carbon atoms with 2n (16) Isomers. n = unequal number of carbon atoms.
Epimers: a sugar that differs only in the configuration around a single carbon atom. E.g. D-glucose and D-Mannose 2. D-Glucose and D-Galactose
Diastereomers: One type of diastereomers (or geometric stereoisomers) differs in terms of “cis” and “trans”. In diastereomers, some chiral centres are similar and some are opposite. The molecule does not resemble a mirror image of its diastereomer.
D and L isomers:
Enantiomers: Enantiomers are mirror image molecules that cannot be elevated to each other. The suggestion suggests that two mirror molecules can be psychologically integrated into one object as they are integrated. Eg, D-glucose and L-glucose. When the OH group in the carbon atom adjacent to the terminal primary alcohol (carbon atom 5 in the right), sugar is a member of series D. On the left is a member of the L series. Most monosaccharaides are classified as D.
Optical Isomerism: When a cooled light beam is transferred to a solution that reflects light performance, the part will be moved right or left depending on the type of combination present. The element that alternates the illuminated light to the right is said to be dextrorotatory and the plus (+) sign is used for designation. The rotation of the pole on the left (laevorotatory action) is marked with a minus sign (-). When an equal number of dextrorotatory and laevorotatory isomers are present, the resulting mixture has no optical functions, because the functions of each isomer overlap. That mixture is said to be a racemic mixture.
.
There are three categories of carbohydrates:
1. Monosaccharides
2. Oligosaccharides
3. Polysaccharides
Monosaccharides or simple sugars:
It contains one unit of polyhydroxy aldehyde or ketone. E.g.: Sugar, Fructose, Galactose.
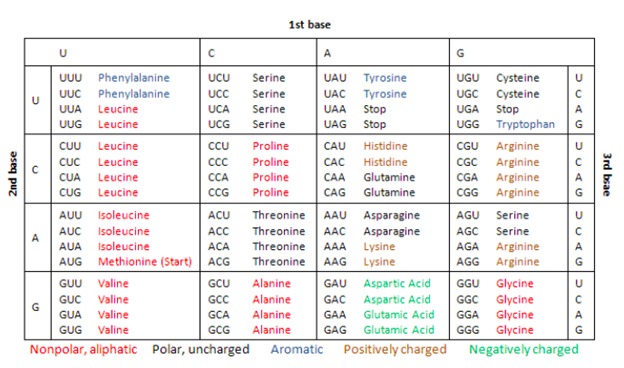
Once the group is at the end of the carbon chain (i.e., in the aldehyde group) monosaccharide is an aldose. If the group can be in another position (in the ketone group) monosaccharide is ketoses. The simplest monosaccharaides are trios-carbon trioses including Glyceraldehyde aldotriose, and Dihydroxyacetone ketotriose.
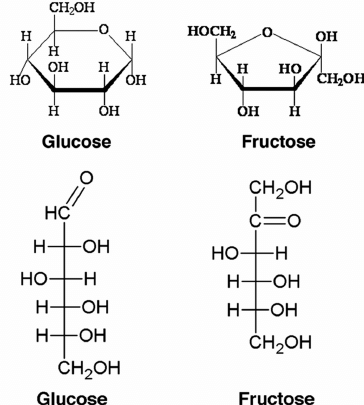
Glucose, a major source of energy for many living things. Glucose is present in both open chains and a ring form with rings forms when glucose is dissolved in water. It contains radical aldehyde as part of the structure. Group C = O in the aldehydic group has reduced concentrations and therefore lowers blood sugar. This is the most prominent monosaccharide in natural D-glucose, sometimes called dextrose. Fructose has a ketonic group as part of the structure. Group C = O in the group has reduced areas which is why it reduces sugar. Galactose is an aldohexose with reduced properties. One of the ingredients of Lactose.
Oligosaccharides: composed of short chains of monosaccharide units bound by glycosidic bonds. The most abundant are disaccharides, which have two monosaccharide units connected by glycosidic bonds. E.g., Maltose, Lactose, Sucrose.
• Maltose is also known as a sugar source made up of two sugar molecules. Maltose is formed by joining two alpha glucose molecules that meet the condensation reaction and form a glycosidic bond between molecules. It has reduced properties.
• Sucrose is a common household sugar or sugar cane and is composed of monosaccharides glucose and fructose bond. Sucrose does not contain an unknown free carbon atom.Anomeric carbons atoms of both monosaccharide units form a glycosidic bond. So sucrose sugar does not decrease. Non-reducing disaccharides are called glycosides.
Polysaccharides: Sugar polymers contain more than 20 or more units of monosaccharides called polysaccharides. E.g., Starch (Amylose, Amylopectin), Glycogen, Cellulose, Chitin. Polysaccharides are the main polymers of monosaccharaides .The polysaccharides may not be soluble or form colloidal suspensions. Starch is an alpha glucose polymer that is a mixture of two different polysaccharides.
AMYLOSE AND AMYLOPECTIN
AMYLOSE: They are long, unstable plates of sugar units. It is made up of a series of condensation reactions that include alpha glucose molecules that have been synthesized into an extended chain that forms many glycosidic bonds
AMYLOPECTIN – a very powerful polymer for glucose units. It contains an open series of alpha glucose units with branch points across all twelfth glucose. Branch points are formed when carbon 6 of the glucose molecule within an open chain forms a glycosidic bond with carbon 1 of the glucose molecule placed above the series

Glycogen
• Glycogen is often referred to as glycogen. The structure of Glycogen is almost identical to amylopectin but there are many branches in glycogen. Glucose is stored as glycogen in large mountains in both liver and bone tissue.
Cellulose:
• Cellulose is one of the most important structural polysaccharides because it is the major component of plant cell walls. Many identical chains of beta glucose units are formed and the whole chain contains hydrogen bonds between groups of OH adjacent chains.
Chitin:
• Chitin can be a polysaccharide that makes many invertebrate exoskeletons. N-acetyl glucosamine polymer in beta 1 to 4 glycosidic bonding. It is a key component of the insect and crustacean sac that protects and supports.