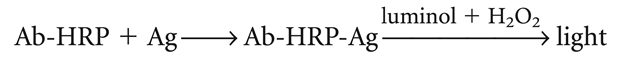BY- SAI MANOGNA (MSIWM014)
Radioimmunoassay (RIA):
Radioimmunoassay (RIA) is one of the most responsive antigen or antibody detection techniques. In 1960, this procedure was first developed by two endocrinologists A. Berson and Rosalyn Yalow, to evaluate levels in diabetics of insulin-anti-insulin complexes. While their technique addressed some skepticism, at concentrations of 0.001 micrograms per millilitre or less, it soon proved its usefulness for testing hormones, serum proteins, medicines, and vitamins. The importance of the technique was recognised in 1977, some years after Berson ‘s death, by the granting of the Nobel Prize to Yalow.
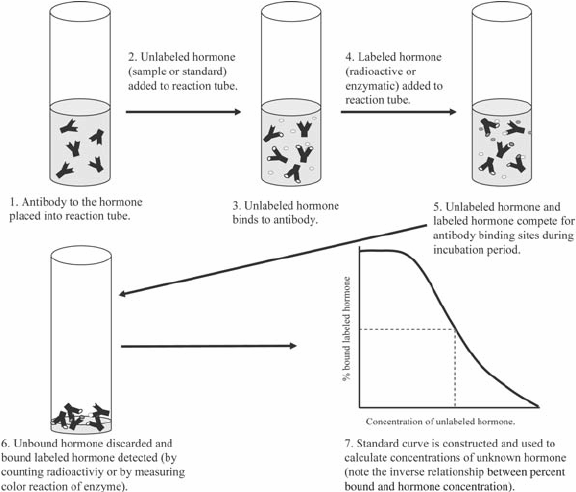
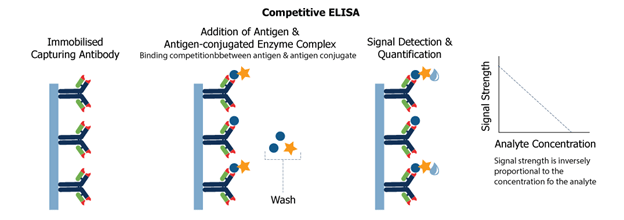
Principle : RIA’s technique involves the competitive binding to a high-affinity antibody of radiolabeled antigen and unlabeled antigen. The labeled antigen is mixed with an antibody at a concentration that saturates the antigen binding sites of the antibody.. Then, in increasingly greater quantities, test samples of unlabeled antigen of unknown concentration are added. The antibody does not differentiate between labelled and unlabeled antigen, so the two kinds of antigen compete against the antibody for available binding sites. If the concentration of unlabeled antigen increases, it will displace more labelled antigen from the binding sites. In order to assess the amount of antigen present in the test sample, the decrease in the quantity of radiolabeled antigen bound to a particular antibody in the presence of the test sample is measured.
Procedure :
A gamma-emitting isotope such as 125I is commonly labelled with the antigen, but beta-emitting isotopes such as tritium (3H) are often used as labels. The radiolabeled antigen is part of the assay mixture; a complex mixture, such as serum or other body fluids, containing the unlabeled antigen (test sample).
1. The first step in setting up an RIA is to decide the amount of antibody required in the assay mixture to bind 50 percent to 70 percent of a fixed amount of radioactive antigen.
2. Unlabeled antigen applied to the sample mixture would also compete for the limited supply of antibodies with radiolabeled antigen. (Even a small amount of unlabeled antigen added to the labelled antigen and antibody assay mixture will cause the amount of radioactive antigen bound to decrease, and this decrease will be proportional to the amount of unlabeled antigen added).
3. The Ag-Ab complex is precipitated to distinguish it from free antigen (antigen not bound to Ab) to assess the quantity of labelled antigen bound, and the radioactivity in the precipitate is calculated.
4. Using unlabeled antigen samples of a known concentration (in place of the test sample), a standard curve can be produced and the amount of antigen in the test mixture can be accurately calculated from this map.

For the separation of the bound antigen from the free antigen in RIA, several methods have been established. One strategy involves precipitating the complex of Ag-Ab with a secondary antiserum anti-isotype.
For example, if rabbit IgG antibodies are included in the Ag-Ab complex, then goat anti-rabbit IgG will bind to rabbit IgG and precipitate the complex.
Another technique makes use of the fact that Staphylococcus aureus protein A has a high IgG affinity. If the Ag-Ab complex produces an IgG antibody, it can be combined with formalin-killed S to precipitate the complex. The amount of free labeled antigen remaining in the supernatant can be determined in a radiation counter after removal of the complex by one of these methods; subtracting this value from the total amount of labelled antigen added yields the amount of labeled antigen bound.

Advantages:
- Used to detect very small amounts of serum antigens and antibodies.
- Used to quantify hormones, pharmaceutical products, HBsAg and other viral antigens.
- Analysis of nanomolar and picomolar concentrations in biological fluids.
Disadvantages:
- Cost effective
- Radio-labeled compounds have short shelf life
- The concerns surrounding the handling of radioactive (nuclear) wastes.