BY: ABHISHEKA G (MSIWM013)
INTRODUCTION:
Among all the advanced techniques in the field of biotechnology gene therapy is one of the important milestones. Gene therapy is a technique used in biotechnology in which new foreign genes are inserted into existing cells of living organisms to treat diseases. This technique is employed to replace the defective genes in the genome which causes diseases. The research on gene therapy was first started in 1972 by Friedmann and Roblin who published a paper entitled ‘gene therapy to cure genetic disorders. Later in the future, the research was developed by scientists, and on the 14th of September 1990, for the first time the gene therapy clinical trials were approved by the National Institute of Health (NIH), the US under the guidance of William French Anderson, which was performed on 4 years old girl who was suffering from severe combined immunodeficiency.
TYPES OF GENE THERAPY:
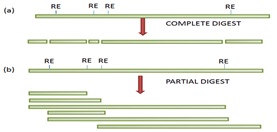
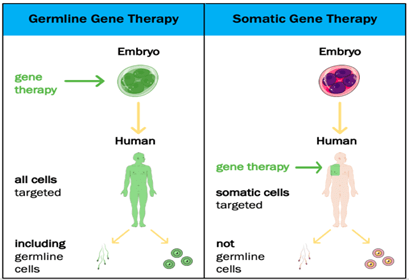
The technique gene therapy is classified into two types based on cells in which foreign gene is inserted, a) Germline gene therapy
b) Somatic cell gene therapy
Germline gene therapy: In this type, the foreign gene or therapeutic gene is inserted into the germ cells of the living organism. In this technique the inserted genes are passed from generation to generation, hence they are heritable.
Example: The therapeutic cells are inserted into the germ cells like sperm and egg.
Somatic cell gene therapy: In this technique, the foreign gene or therapeutic gene is inserted into the somatic cells of the living organism. In this technique, the inserted genes do not pass from generation to generation.
Example: Insertion of genes into bone marrow, blood cells, skin cells, etc.
APPROACHES OF GENE THERAPY:
There are two ways of approaches are available in gene therapy:
a) In-vivo gene therapy
b) Ex-vivo gene therapy
In-vivo gene therapy: In this technique, the target cell is fixed in the body of the living organism and the therapeutic gene or foreign gene is inserted into the target cell through a vector. The vector used to insert the therapeutic gene into the target cell is either a viral or non-viral vector. This technique is mainly used to treat genetic disorders.
Example: It is used to treat the genetic disorder cystic fibrosis.
Ex-vivo gene therapy: In this technique, the therapeutic genes are inserted into the cultured cells in the laboratory.
Procedure: The genetically defective cells are isolated from the living organism and they are cultured inside the laboratory. Then the therapeutic genes are inserted into the cultured cells in the laboratory. After insertion, the genetically corrected cells from the cultured cells are isolated and grown in the laboratory, then the modified cells are transplanted into the living organism.
Example: The first gene therapy was performed on 4 years old girl Ashanthi Disilva, who was suffering from severe combined immunodeficiency (SCID), which is caused due to defects in the gene which codes for adenosine deaminase. This is an example of ex-vivo gene therapy.

VECTORS IN GENE THERAPY:
Vector is the vehicle of gene delivery, an agent that carries to the therapeutic or desired gene into the target cell. There are two categories of vectors are employed in gene therapy.
- Viral vectors
- Non-viral vectors
Viral vectors: The viruses are used as vectors to deliver the desired gene to the target cell of the living host.
Example: Retrovirus: The genetic material of the virus RNA and it can carry a DNA of size less than 3.4kb and integrated it into the host genome of stable fashion, and it targets the cells in dividing the state.
Adenovirus: The genetic material of this virus is double-stranded DNA, they carry and integrate the desired DNA into target cells that are a non-dividing state.

Non-viral vectors: Some of the chemicals and physical instruments are used to carry desired DNA into the host cell are called Non-viral vectors.
Some of the physical methods to insert the desired DNA into the target cell are Electroporation, gene gun, etc.
Electroporation: In this technique, the electric shock of small pulses of high voltage is used to carry the DNA across the cell membrane of the target cell. After the small electric shock, the temporary pores are caused in the cell membrane to provide the passage for desired DNA into the host cell.
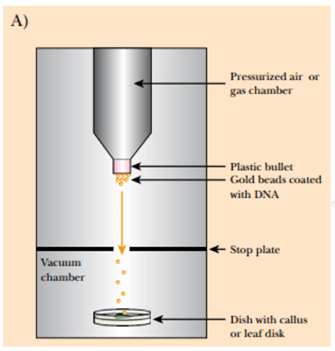
Gene gun: In this method, the gold particles or tungsten particles are coated with the desired DNA molecule, then the coated DNA is placed into a device called Gene gun which generates the high pressure necessary for the penetration of the desired DNA into the target or host cell and the gold and tungsten particles are left behind on stopping disc.
Sonoporation: In this method, the DNA is inserted into the target cell with help of ultrasonic frequencies.
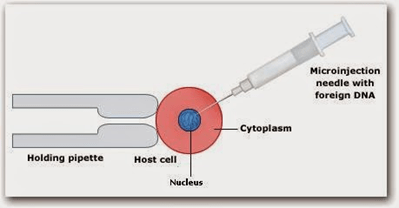
Microinjection: In this method, microscopic materials are inserted into the target cell with the help of a small microscopic device called Micromanipulator.
Some of the chemical methods used to carry desired into the target cell are lipofection and using detergent mixtures.
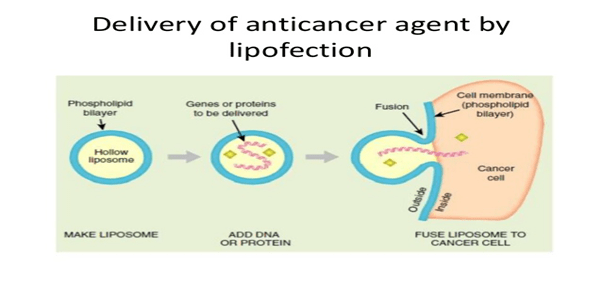
Lipofection: In this technique, the liposomes are used as a vector to insert the desired DNA into the target cell. The liposomes are artificial phospholipid vesicles used to transport various molecules into cells including DNA.
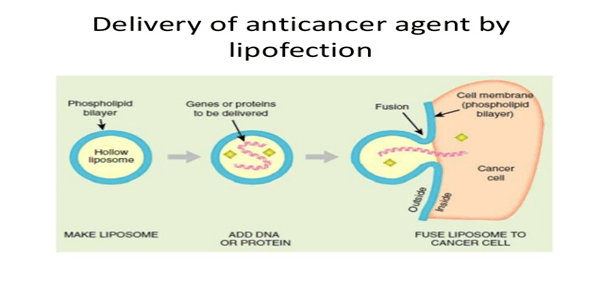
Detergent mixtures: Some of the detergent mixtures like calcium phosphate are mixed with cDNA molecules and they are inserted near to the target cells. Then these detergent mixtures disturb the cell membrane od the cell and increase the size of the pore through which the cDNA enters into the cell.
ADVANTAGES OF GENE THERAPY:
- Gene therapy plays an important role to replace the defective disease-causing genes in the genome.
- It also helps treat genetic disorders and some of the deadly diseases like cancer.
DISADVANTAGES:
- Diseases caused by defects in multiple genes are not treated by gene therapy.
- Some of the viral vectors cause toxicity and inflammatory reactions.
- Some of the ethical issues arise.