BY: RAHUL ANDHARIA (MSIWM001)
Hypersensitivity:
It refers to excessive, undesirable damage posing, discomfort producing and sometimes fatal reactions produced by the body’s own natural immune system. In simpler terms it simply refers to allergic reactions caused by immune system.
Introduction:
- During hypersensitivity, the host is in pre-sensitized immune state. This simply means that the host was primarily exposed once to the antigen, and comes in contact again for the second time with the same antigen.
- Based on the mechanisms involved and time taken for allergic reaction, hypersensitivity reactions are classified into four types: Type I, Type II, Type III, and Type IV hypersensitivity reactions.

Type I Hypersensitivity:
It is also known as immediate or anaphylactic or atopic hypersensitivity. Type I hypersensitivity is generally caused due to reexposure to the specific type of antigen or allergen(substance which causes allergy). It can lead to systemic or local reactions and involves eyes(conjunctivitis), skin(eczema), nasopharynx(rhinitis) and gastrointestinal tracts(gastroenteritis). Symptoms generally varies from mild to even fatal and can lead to death or anaphylactic shock. The time of reaction ranges from 10-15 minutes after the action of allergen, but sometimes there is delayed onset(10-12 hours).
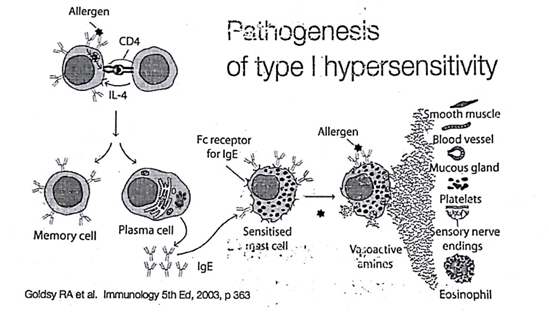
Mechanism:
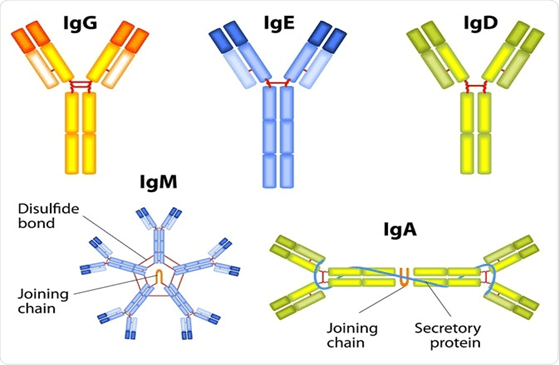
- The reaction is mediated by IgE antibody. Primary cellular component involved is mast cell or basophil cells.
- Upon very first exposure to allergen, APC(antigen presenting cells) processes the antigen and present it to Th2 cells.
- Upon released of IL-4 and IL-12 by Th2 cells, B cells gets activated.
- After that, the B cells differentiate into plasma cells. Plasma cells synthesise and secretes IgE antibody (type of antibody usually involved in allergic reactions).
- IgE binds to FC(fragment crystallization) of mast cells and sensitize it.
- Upon subsequent exposures to the same antigen, now the mast cells bind to IgE antibody and releases the inflammatory molecule. This will result in allergic symptoms.
Diagnosis:
- Involves skin tests( pricks and intradermal samples)
- Specific and total IgE antibody measurements against the suspected allergen by using ELISA(Enzyme linked immunosorbant assay).
- Increased level of IgE in the ELISA test, will confirm allergy or atopic condition.
Treatment:
- Anti-histamines for symptomatic treatment. Anti-histamines can block the histamine receptors.
- Cromo Lyn sodium, a chemical can inhibit mast cell degranulation. It does it by inhibiting ca+2 influx.
- IgG antibodies can be used as it can bind with IgEs FC portion and prevents mast cell sensitization.
Examples of type I hypersensitivity: Conjunctivitis, Eczema, Eosinophilia, Angioedema.
Type II Hypersensitivity:
This type II hypersensitivity is also known as antibody dependant or cytotoxic hypersensitivity and is known to affect many organs and tissues. The antigens are normally endogenous. However, some exogenous chemicals or molecules such as haptens(substance which combined with proteins can produce antibodies) can also induce type II hypersensitivity. Reaction time is usually from few minutes to hours.
Mechanism:
- IgG and IgM antibodies bind to the antigen and form complexes. This complexes than results in activation of compliment pathway(pathway through which foreign antigens are destroyed).
- At the site of membrane attack complexes, mediators of acute inflammation are generated, which will lead to cell death and lysis.
- Phagocytes, expressing FC receptors can induce phagocytosis.
- The FC receptors, recognises surface bound antibody and compliment proteins .
- ADCC(antigen dependant cell mediated cytotoxicity) is another form of hypersensitivity where tags of IgG and IgM antibodies bind. Macrophages and NK(natural killer cells) than recognises this tags and kills them.
Diagnosis:
- It includes detection of circulated antibodies against the tissues involved.
- Detection of presence of antibodies in the liquid biopsy through immunofluorescence.
Treatment:
- Anti-inflammatory agents
- Immunosuppressing agents
Examples of type II hypersensitivity: Erythroblastosis Fetalis, Hashimotos thyroiditis, transfusion reactions, Rheumatic fever.

Type III Hypersensitivity:
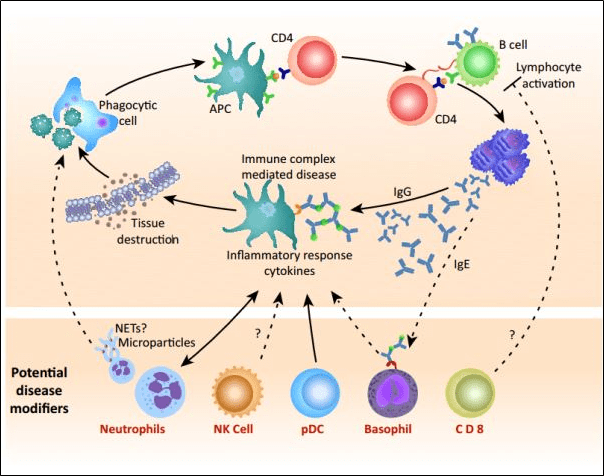
Also called as immune complex hypersensitivity. The reaction is general or may involve organs such as skin, blood vessels, kidneys and joints. This reaction is caused primarily by micro-organisms. The reaction time is 3-10 hours after being exposed to the allergen.
Mechanism:
- Mediated by soluble immune complexes. Mostly the complexes are of IgG class but sometimes they can be of IgM class.
- Exogenous(viral or chronic infections) or endogenous (non specific autoimmunity) antigens.
- Primary components are compliment proteins( C3a, 4a and 5a).
- Major damage is caused by platelets and neutrophils.
- Macrophages and neutrophils are present in the lesions.
- Antibody deposition triggers an immune response according to classical compliment pathway. There is formation of 2complexes(complex is formed when IgG and IgM are bounded to antigen)The larger complex gets eliminated but the first complex remains and the antigen antibody complex will spread and deposit.
Diagnosis:
- Examinationof tissue biopsies for immunoglobulins and compliment by immunofluorescence microscopy.
- Raji cell test and polyethylene glycol tests can be performed to measure immune complexes.
Treatment:
- Anti-inflammatory agents.
Examples of type III Hypersensitivity: Arthur’sreaction, Rheumatoid arthritis, symptoms of malaria.

Type IV Hypersensitivity:
Also can be called as delayed type or cell mediated hypersensitivity. Classic example of this type of hypersensitivity is tuberculin reaction(montoux) which peas after 48 hours of antigen injection. Erythema and induration(thickening of skin) are the characteristics of the lesion.
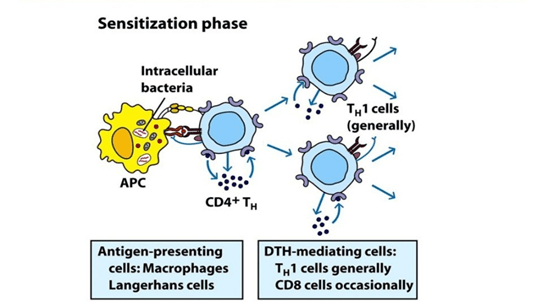
Mechanism:
- Antigen in the complex is recognised by cytotoxic Cd8 T cells and Cd4 helper T cells by MHC I or MHC II complex.
- IL-1 secreted by macrophages further helps in proliferation of Cd4 T cells.
- Th1 mediated response is elucidated upon reexposure to antigen.
- Cd4 T cells secretes IL-2 which further induces the release of other type 1 cytokines and thus induce immune response.
- Cd8 cells when activated, destroys target cells upon contact. Macrophages produces hydrolytic enzymes and when comes in contact with intracellular pathogens, macrophages transforms into multinucleated giant cells.
Diagnosis:
- Montoux test and patch test in-vivo is generally performed.
- Some of the in-vivo tests for delayed hypersensitivity can be mitogenic response, nephro-cytotoxicity, and production of IL-2.
Treatment:
- Corticosteroids
- Immunosuppressive agents
Examples of type IV Hypersensitivity: symptoms of tuberculosis, coeliac disease, symptoms of leprosy.
Type V Hypersensitivity:
There is an additional type of hypersensitivity that is, type V(5). This usually occurs when the IgG antibodies have a effect towards their target. In this, the antibody binds to the cell surface receptors rather than binding to cell surface components and as a result it prevents the binding of the intended ligand to the receptor and thus impairs cell signalling.
Example of type V: Graves disease.

- Diagram showing all four hypersensitivity reactions.